Anthropology
Anthropology at Kent 1965-2026
The subject of Anthropology was taught at the University of Kent from 1965-2026. Paul Stirling was appointed Professor of Sociology in 1965 and established both Sociology and Social Anthropology at the university.
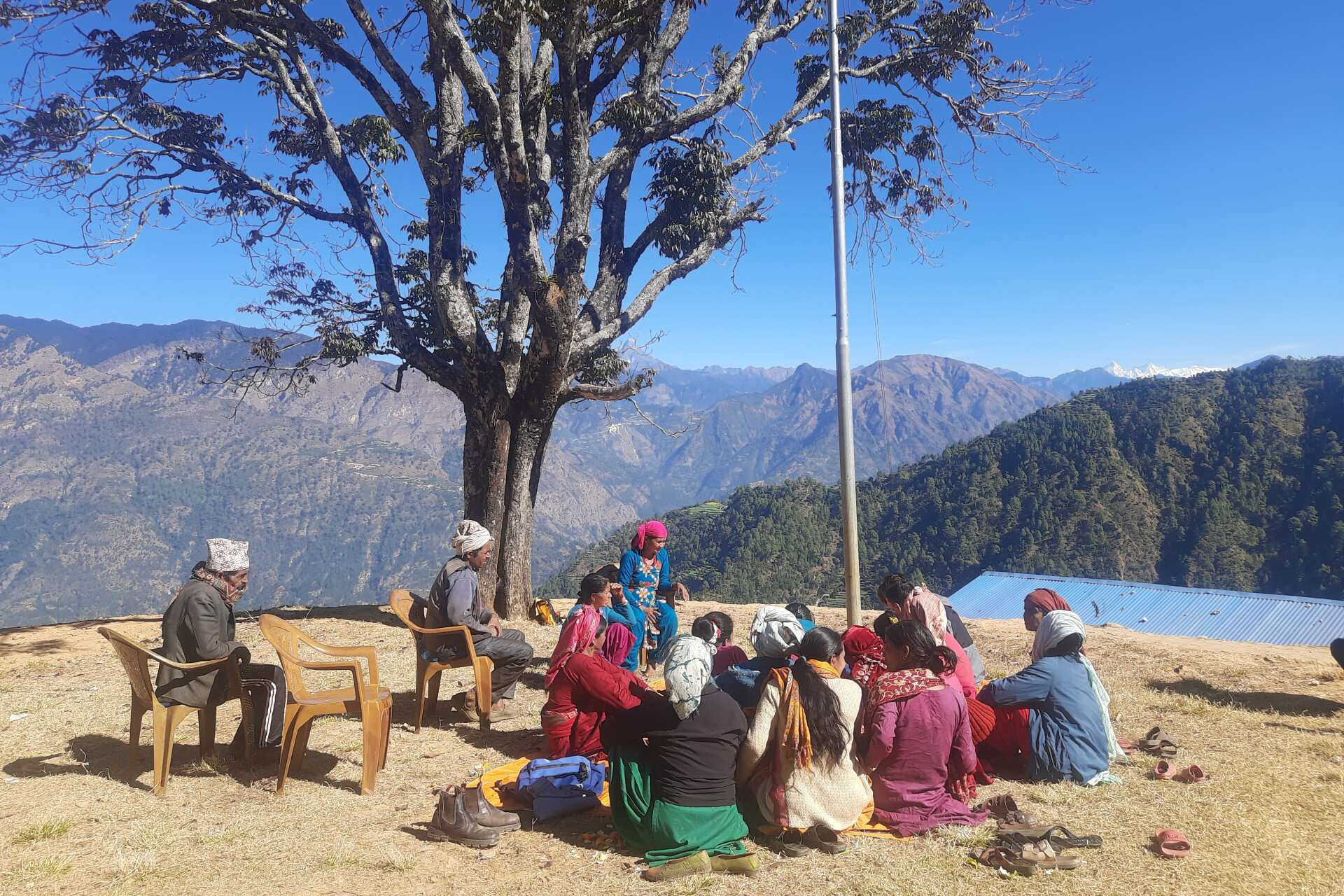
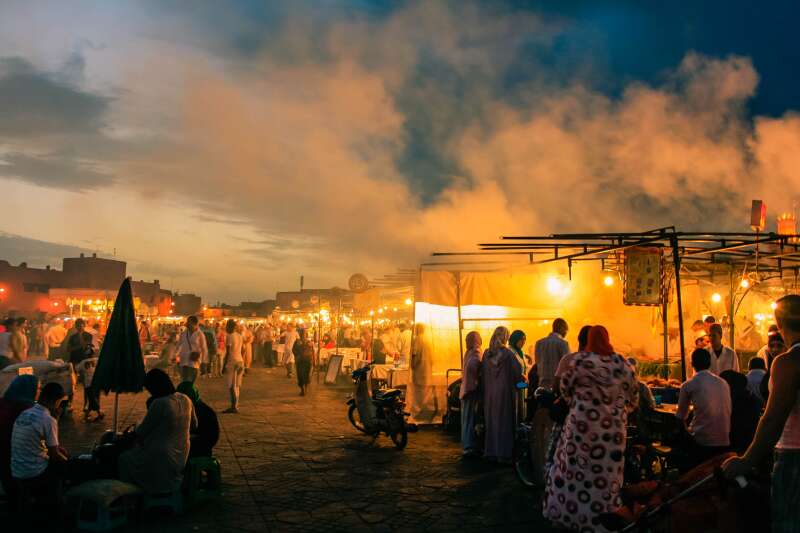
Since 1965 the University of Kent has employed over 100 anthropologists, including 15 full professors. Anthropology at Kent began with a specialist focus on the Mediterranean and earned an international reputation as a centre of research excellence in Mediterranean studies.
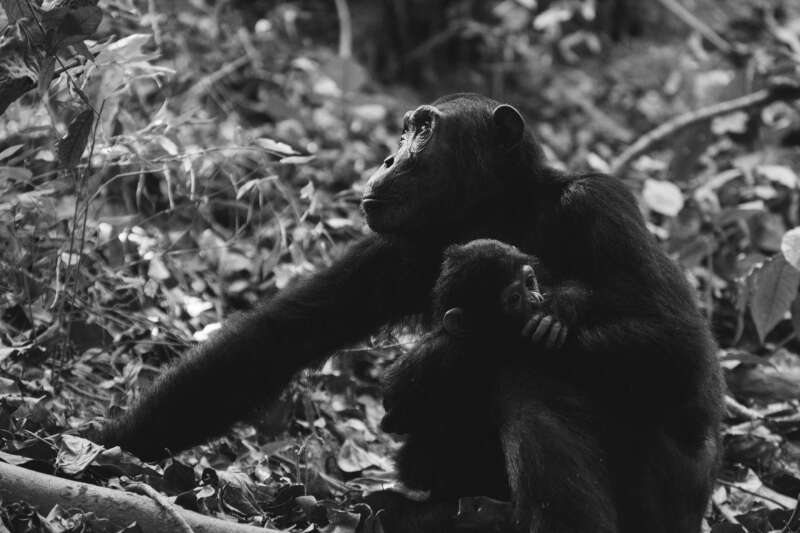
As Anthropology at Kent grew, it developed new areas of excellence, including social anthropology and computing, ethnobiology, and biological anthropology. There are over 1550 anthropology publications in the Kent Academic Repository.In the late 1990s Anthropology split from Sociology and allied with the Durrell Institute of Conservation and Ecology, eventually forming the School of Anthropology and Conservation in 2010.
The School was closed in 2024, along with all undergraduate and postgraduate anthropology courses. Over 60 years Kent Anthropology produced over 160 PhDs and trained thousands of undergraduate and taught postgraduate students.
The University of Kent is still home to several anthropology colleagues working in other schools and a thriving community of honorary and emeritus anthropologists.
In addition to anthropologists working in other schools, we’re proud to have honorary and emeritus anthropology staff members who continue to share their knowledge and experience.
Whether you’re a student, researcher, or community member, we are happy to offer guidance, answer questions, and connect with you.
Showing profiles
Filters
Filter by profile name or research interest.
Research
We engage with local, national and international partners to produce high-quality research that has a positive impact in the wider community. By combining laboratory research with fieldwork, academic staff and students are able to monitor and survey in depth a variety of topics, ranging from ethnographic studies to forensic bio archaeology, species conservation and land-use changes.


